February 15, 2021 -- Researchers have identified interactions between short viral proteins and receptors that facilitate the entry of the SARS-CoV-2 virus into a host. This evidence of molecular links, published in Science Signaling on February 12, may help scientists identify drugs that are highly effective at blocking the virus.
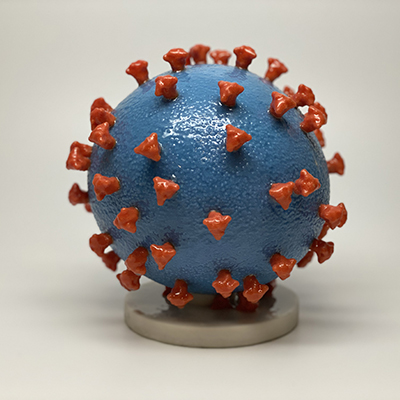
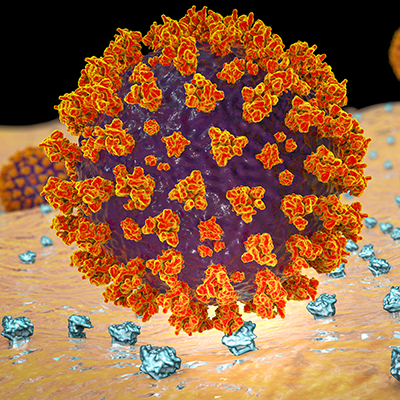
SARS-CoV-2 uses the angiotensin-converting enzyme 2 (ACE2) as a receptor to attach to host cells. ACE2 is a single-pass type I membrane protein with a short cytosolic C-terminal region. While SARS-CoV-2 primarily binds to ACE2 in the lungs, previous studies have revealed that there is very little ACE2 gene expression in normal lungs, suggesting that the ACE2 receptor is insufficient on its own to establish severe lung disease.
The current study, led by researchers at the European Molecular Biology Laboratory (EMBL) and in collaboration with teams from the Universidad Nacional de San Martín in Buenos Aires, Merck, and the University College Dublin, suggests that receptors, such as integrins that bind a wide variety of ligands with an RGD (arginine-glycine-aspartate) sequence motif, could act as co-receptors for cell attachment.
Integrins are cell attachment receptors that are known to be targeted by a range of viruses for cell entry and activation of intracellular pathways. They are unique in the fact that they stimulate signals bidirectionally -- extracellular ligands can induce cytoplasmic pathway activation, but also intracellular interactions with cytosolic tails can affect ligand-binding affinity. This complexity is due to the dimeric structure of integrins which are composed of two subunits.
Some viral proteins contain RGD short linear motifs (SLiMs) that can modulate integrin activation. Some viruses such as HIV can even incorporate integrins into their own membranes to mediate interactions with host cells. Viruses hijack host processes by using SLiMs to establish protein-protein interactions. Acting as molecular switches, integrins through phosphorylation can affect cell states to facilitate viral entry.
"SLiMs could 'switch' to turn viral entry signals on or off," said senior author Lucía Chemes, PhD, researcher at Universidad Nacional de San Martín, in a statement. "This means that if we can find a way to reverse these switches using drugs, this might stop coronavirus from entering cells."
Many viruses use endocytosis to enter host cells. Receptor-mediated endocytosis (RME) is a cellular import process triggered by cell surface receptor proteins, including any cargoes attached to them, in which vesicles are assembled entirely through cooperative low-affinity interactions of SLiMs and phospholipid head groups with their globular protein domain partners. SARS-CoV-2 enters host cells through protease-mediated activation of the spike protein to facilitate viral fusion, utilizing endocytosis.
Coronaviruses also exploit the autophagy (catabolism of cell components) machinery through different mechanisms. For instance, SARS-CoV-2 can confine double-stranded RNA within double-membrane vesicles to conceal the viral genome from the innate immune system.
"If SARS-CoV-2 targets proteins involved in endocytosis and autophagy, it means these processes might be hijacked by the virus during infection," said first author, Bálint Mészáros, PhD, a postdoctoral in the EMBL team.
Finding SLiMs
The researchers identified a set of conserved SLiM candidates in ACE2 and integrin proteins, that are likely to act in the cell entry system of SARS-CoV-2. The eukaryotic linear motif (ELM) resource, a database of over 280 manually curated SLiM classes with experimental evidence, was used to quickly identify and compare candidates.
They tested the ability of SLiMs to bind partner protein domains with an expected low or high affinity in vitro. The ACE2 basement membrane (PBM), the Src family kinases (SFK) Src Homology 2 (SH2)-binding motif, the adapter protein (AP) µ2-binding motif, as well as the integrin β3 phospho-LC3-interacting region (LIR) were identified for their potential roles in cellular endocytic-autophagosome pathways during SARS-CoV-2 infection.
These sequences may help scientists understand how the virus recognizes target membranes, enters into cells, and repurposes membrane components to drive its own replication.
Drugs that count SLiM action
The researchers' analysis suggested that SARS-CoV-2 hijacks both ACE2 and integrins, co-opting SLiMs to drive viral attachment, entry, and replication. Host directed therapies can be used to prevent viral entry. The team then gathered a list of existing drugs that interfere with endocytosis and autophagy that may be effective at blocking SARS-CoV-2 infection.
They identified several candidate drugs that could interfere with SLiM-induced endocytosis or autophagy. For instance, cilengitide, a selective integrin inhibitor, may be useful in blocking virus attachment to target cells, but also in RGD-dependent binding of pathogens that protects against sepsis. The antibody abituzumab (DI-17E6) is a pan-α integrin antibody that has been shown to have potential for blocking virus entry. Furthermore, integrin inhibitor, GSK3008348, has been shown to have an effect on lung fibrosis in a mouse therapeutic model.
"If clinical trials prove some of these drugs to work against COVID-19, this could be a game changer," explained senior author, Manjeet Kumar, PhD, a bioinformatics scientist on the EMBL team.
Copyright © 2021 scienceboard.net