June 2, 2020 -- A new study has revealed that SARS-CoV-2 infects the nasal cavity to the greatest degree and infects and replicates progressively less well in cells lower in the respiratory tract. The work, including the development of key laboratory tools, has been accepted by Cell.
The SARS-CoV-2 genome contains 16 nonstructural proteins, four structural proteins (spike, membrane, envelop, and nucleocapsid), and eight accessory proteins. The spike protein mediates viral entry by binding to the human angiotensin-converting enzyme 2 (ACE2) followed by proteolytic processing by TMPRSS2 and furin. Therefore, spike proteins are the main target of host neutralizing antibodies.
SARS-CoV-2 primarily infects cells in the respiratory tract and causes a broader range of symptoms than its predecessors MERS-CoV and SARS-CoV. To further understand the organ-level architecture of receptor expression, researchers at the University of North Carolina at Chapel Hill (UNC) School of Medicine and Gillings School of Global Public Health developed a combination of RNA in situ hybridization techniques and novel SARS-CoV-2 reporter viruses by reverse genetics.
Reverse genetics allowed the researchers to generate a green fluorescent protein (GFP) reporter virus to explore SARS-CoV-2 pathogenesis and a luciferase reporter virus to demonstrate that sera from SARS and COVID-19 patients exhibited limited cross-coronavirus neutralization.
To evaluate viral RNA synthesis, the researchers used northern blot analyses to differentiate between the reporter viral samples and clinical isolates and wildtype samples.
"This is a landmark study that reveals new and unexpected insights into the mechanisms that regulate disease progression and severity following SARS-CoV-2 infection," said Ralph Baric, PhD, the William R. Kenan Distinguished Professor of Epidemiology at the UNC Gillings School of Public Health, in a statement. "In addition, we describe a new reverse genetic platform for SARS-CoV-2 allowing us to produce key indicator viruses that will support national vaccine efforts designed to control the spread and severity of this terrible disease."
The researchers found that serine proteases like TMPRSS2 and furin enhance the replication efficiency of SARS-CoV-2 in vitro. They also confirmed that previously described neutralizing antibodies against MERS-CoV and SARS-CoV did not neutralize SARS-CoV-2, even though SARS-CoV-2 shares around 75% sequence identity with SARS-CoV.
As part of the study, the researchers also characterized ACE2 expression in the upper airways (nose, oropharynx) and lung (lower airways, alveoli) using high-sensitivity RNA in situ hybridization. The analysis found the highest ACE2 expression in the nose and decreasing expression throughout the lower respiratory tract. In parallel, there was a gradient of SARS-CoV-2 infection from the upper respiratory tract to the lower respiratory tract. Most of the ACE2-expressing cells were ciliated cells, not secretory or goblet cells.
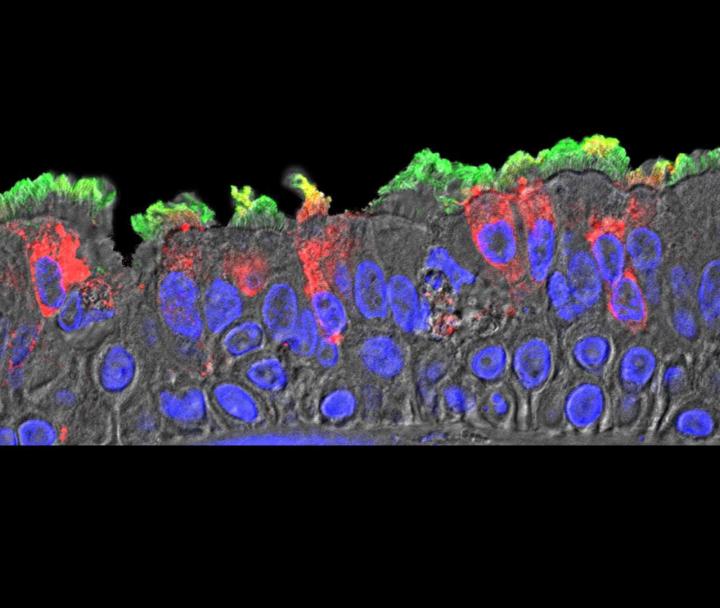
The researchers offered that nasal surfaces may be the dominant initial site for SARS-CoV-2 respiratory tract infection, primarily composed of ciliated cells in the superficial epithelium. Oral-lung aspiration (drawing breath) may contribute to spreading of the infection to the lower airway. The results of COVID-19 autopsy studies confirm that patchiness of SARS-CoV-2 infection matches the variable nature and pathogenesis of an infection that originated from higher in the airway.
The researchers explained that techniques developed from the study, including a version of SARS-CoV-2 reengineered to carry a fluorescent beacon, will be useful in future investigations of the virus.
"These results, using some novel and innovative methodology, open new directions for future studies on SARS-CoV-2 that may guide therapeutic development and practices for reducing transmission and severity of COVID-19," said James Kiley, PhD, director of the Division of Lung Diseases at the National Heart, Lung, and Blood Institute, part of the National Institutes of Health.
Do you have a unique perspective on your research related to virology or infectious disease? Contact the editor today to learn more.
Copyright © 2020 scienceboard.net