April 23, 2021 -- Researchers have identified naturally occurring molecules that are created from the breakdown of hemoglobin and block the binding of a subset of human antibodies to SARS-CoV-2. The discovery, published in Science Advances on April 22, may help explain why some COVID-19 patients can become severely ill despite having high levels of antibodies against the virus.
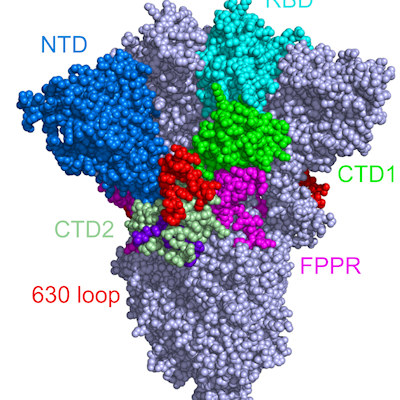
The SARS-CoV-2 spike protein is made up of two subunits, S1 and S2, which mediate binding to the receptor and facilitate fusion. The recognition of SARS-CoV-2 host receptor, the cellular membrane protein angiotensin-converting enzyme 2 (ACE2), maps to the S1 C-terminal domain (referred to as the receptor-binding domain [RBD]).
Most neutralizing antibodies target the RBD. Meanwhile, the function of the N-terminal domain (NTD) is relatively unknown and mutations in the NTD are associated with viral escape from antibody immunity.
While conducting COVID-19 research, teams from the Francis Crick Institute, in collaboration with researchers at Imperial College London, King's College London, and University College London identified that the SARS-CoV-2 spike and S1 protein carried a distinctive green hue, which was confined to the NTD and absent from the RBD. They isolated the pigment and confirmed the presence of biliverdin by mass spectrometry.
The production of biliverdin from the heme moiety of hemoglobin in the blood is the first major breakdown step, after which the enzyme biliverdin reductase performs the second step, producing bilirubin. Elevated bilirubin levels have been correlated with the severity of symptoms and mortality among COVID-19 patients. The researchers found that the NTD of the SARS-CoV-2 spike protein strongly binds to biliverdin, giving the proteins an unusual green coloration.
Using cryo-electron microscopy and x-ray crystallography, the team explored interactions between the spike protein and biliverdin. The images revealed that a biliverdin molecule can be buried within a deep cleft on one side of each of the NTD domains. This binding stabilizes the spike protein so that it is not able to open and expose certain parts of its structure.
"When SARS-CoV-2 infects a patient's lungs it damages blood vessels and causes a rise in the number of immune cells," said Annachiara Rosa, PhD, first author and postdoctoral training fellow in the Chromatin Structure and Mobile DNA Laboratory at the Crick, in a statement. "Both of these effects may contribute to increasing the levels of biliverdin and bilirubin in the surrounding tissues. And with more of these molecules available, the virus has more opportunity to hide from certain antibodies. This is a really striking process, as the virus may be benefiting from a side-effect of the damage it has already caused."
Importantly, the authors noted that biliverdin and some SARS-CoV-2 antibodies share the same binding site on the spike protein. This suggests that some antibodies are not able to access their target sites and subsequently cannot bind to and neutralize the virus.
Therefore, the authors hypothesized that the competitive binding masks the antigenic properties of the spike protein and represents another method of immune evasion by the SARS-CoV-2 virus.
By examining blood sera and antibodies from people who were previously infected with SARS-CoV-2, the scientists found that biliverdin could suppress the binding of human antibodies to the spike protein by as much as 30%-50%, with some antibodies becoming ineffective at neutralizing the virus.
The recruitment of the biliverdin leads to stabilization of the NTD structure and makes the viral envelope glycoprotein refractory to neutralization by a subset of human antibodies, the researchers found. They concluded that biliverdin and bilirubin, which are natural molecules present in the body, can suppress the binding of antibodies to the coronavirus spike.
"We were astonished to discover a new trick the virus uses to avoid antibody recognition," said senior author Peter Cherepanov, PhD, a group leader of the Chromatin Structure and Mobile DNA Laboratory at the Crick. "This is a result of a collaborative effort of several amazing teams working at the Crick and three partner universities, led purely by scientific curiosity."
The researchers suggested that strategies to unmask the NTD region of the spike protein are needed in order to make it more visible and accessible to antibodies. They said that this may help lead to new vaccine designs, namely candidates that lack the ability to bind with biliverdin.
The results also highlight the importance of the small pocket in the NTD for the stimulation of antibody immunity against SARS-CoV-2. This has implications for therapeutic development but may also impair the sensitivity of SARS-CoV-2 immunoassays, the team said.
The researchers will continue their work from various angles, including measuring the levels of biliverdin and other haem metabolites in patients with COVID-19 and also exploring if it is possible to hijack the binding site used by biliverdin to potentially find new ways to target the virus.
Do you have a unique perspective on your research related to virology or infectious diseases? Contact the editor today to learn more.
Copyright © 2021 scienceboard.net