January 20, 2023 -- Researchers at Children's Hospital of Philadelphia (CHOP) have developed a new, more accurate computational tool for long-read RNA sequencing. The tool, called Error Statistics Promoted Evaluator of Splice Site Options (Espresso), described January 20 in Science Advances, may allow for better diagnosis of rare genetic diseases caused by disrupted RNA and the discovery of potential therapeutic targets in disease.
An RNA molecule from a gene can be cut and joined, or spliced, in different ways before being translated into a protein. This alternative splicing process allows a single gene to encode several different proteins and occurs in many biological processes, including when stem cells differentiate. In diseases, however, alternative splicing can be dysregulated. Examining the transcriptome -- all RNA molecules stemming from genes --can help reveal a condition's root causes.
Historically, it has been difficult to "read" entire RNA molecules because they are usually thousands of bases long. Instead, researchers have used short-read RNA sequencing, which breaks RNA molecules up and sequences them into much shorter pieces. Computer programs are then used to reconstruct the full sequences. Short-read RNA sequencing can provide highly accurate sequencing data with a low per-base error rate. Nevertheless, the information it can provide is limited.
More recently available long-read platforms can sequence RNA molecules over 10,000 bases in length. These platforms do not require RNA molecules to be broken up before sequencing, but they have a much higher per-base error rate, making it difficult to determine the validity of previously unknown RNA molecules discovered in rare genetic diseases and cancers. This limitation has hampered its widespread adoption.
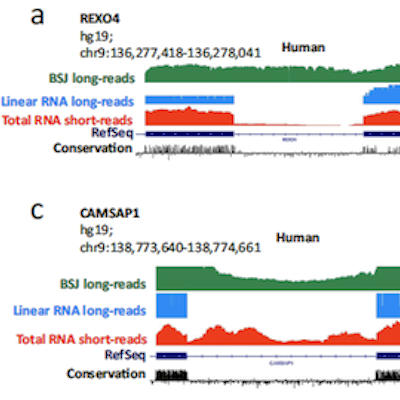
The new computational tool Espresso can more accurately discover and quantify RNA molecules from the same gene -- called RNA isoforms -- using error-prone long-read RNA sequencing data. To do so, Espresso compares all long RNA sequencing reads of a given gene to its corresponding genomic DNA and then uses the error patterns of individual long reads to confidently identify splice junctions, along with their corresponding full-length RNA isoforms. By finding perfect match areas between long RNA sequencing reads and genomic DNA and borrowing information across all long RNA sequencing reads of a gene, the tool can identify highly reliable splice junctions and RNA isoforms, including those not previously documented in existing databases.
The researchers evaluated Espresso's performance using both simulated data and real biological data. They found Espresso performed better than many current tools, both in discovering RNA isoforms and quantifying them. They also generated and analyzed over 1 billion long RNA sequencing reads covering 30 human tissue types and three human cell lines, providing a useful resource for studying human transcriptome variation.
"The transition from short-read to long-read RNA sequencing represents an exciting technological transformation," Yi Xing, PhD, CHOP senior author, said in a statement. "We envision that ESPRESSO will be a useful tool for researchers to explore the RNA repertoire of cells in various biomedical and clinical settings."
Copyright © 2023 scienceboard.net