February 18, 2020 -- Using cryo-electron microscopy (cryo-EM), researchers found unique structures of paramyxoviruses that can lead to a better understanding of viral replication, which in turn can be leveraged to develop improved antiviral drugs. The details were published in the Proceedings of the National Academy of Sciences on February 17.
Paramyxoviruses are enveloped, nonsegmented, negative-strand RNA viruses that cause a wide range of human and animal diseases, including measles, mumps, human parainfluenza, and respiratory syncytial virus. These viruses enter the host by viral attachment, then replicate by making a copy of their negative viral RNA that is complementary to mRNA using RNA polymerase. This copy of viral RNA can then be translated.
The genome of paramyxoviruses encodes several proteins that enable viral replication. The nucleocapsid protein (N) associates with genomic RNA and acts as a template for replication and transcription. Large protein (L) is the catalytic subunit of RNA-dependent RNA polymerase. And phosphoprotein (P) binds the L and N proteins to form the RNA polymerase complex.
The function of these proteins was previously determined by tools like x-ray crystallography and biochemical studies. However, the extent of L-P interface and the specific mechanisms underlying genomic replication and transcription of viral RNA have not yet been determined. To address this concern, Northwestern University researchers and co-lead study authors Robert Lamb, PhD, and Yuan He, PhD, used cryo-EM to solve the structure of the parainfluenza virus 5 L-P complex at 4.3-å resolution.
"Crystallography only works for very orderly and organized proteins," Yuan He said in a statement released by the university. "Virus polymerase complexes are too big to be crystallized and don't have uniformity."
Single-particle cryo-EM allowed the researchers to develop a 3D reconstruction of the proteins. The resulting structure formed a round-shaped globule (five domains within L) with a long tail made of four P proteins and consisted of over 2,000 amino acids.
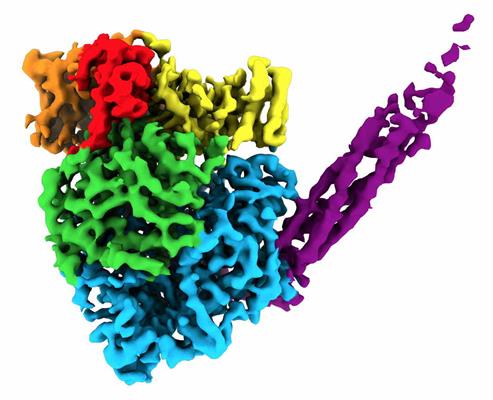
They found that the 3D arrangement of different domains in L is important for RNA synthesis. Specifically, there were two discrete binding interfaces on the L protein surface for binding with the central oligomerization (OD) and C-terminal X domain (XD) domains on the P protein. This binding scheme is necessary for paramyxoviruses to form competent polymerase complexes that work on genomic RNA that is wrapped around the N protein.
Moreover, in comparison to other viruses, the researchers found that large conformational rearrangements occurred of the methyltransferase and C-terminal domain regions within the L protein. This conformational change indicates that the virus uses the same protein to switch between genome replication and transcription.
"This machinery has a dual function," Lamb noted. "It gets both jobs done with one enzyme. The virus's genome is so small, and this gives it economy of scale."
Ultimately, the new insights about viral replication provided by this study could help other researchers design and develop new antiviral drugs for this and other associated viruses -- even coronavirus, which acts similarly to paramyxoviruses.
"This [technique] takes some of the guesswork out of designing drugs," Lamb said. "Traditionally, you have to develop drugs randomly and hope you hit a target, but it doesn't happen very often."
Do you have a unique perspective on your research related to microbiology? Contact the editor today to learn more.
Copyright © 2020 scienceboard.net






