November 24, 2021 -- Scientists have designed bottom-up peptides that can form artificial nanopores to identify and enable single molecule-sorting of genetic material in a lipid membrane, according to new research published in Nature Nanotechnology on November 22.
Protein structures are determined by their polypeptide sequence and give rise to specific protein functionality. This structure depends on the primary sequence of amino acids. Synthetic de novo design of primary sequences of artificial proteins has been an area of research for the past four decades and has the potential to create "molecular machines" that can be used as meaningful targets. In the early days of this field of study, synthetic chemistry and computational design were manual.
Nanopore sensing can be used for label-free, single-molecule detection, where the nanopore-forming membrane protein embedded within a lipid bilayer can form a nanosized pore by allowing target molecules to pass through electrophoretically under applied voltage. Many applications of this technology have been conceived, including low-cost DNA sequencing, small-molecule detection using an adapter or DNA aptamer, nanopore mass spectroscopy, decoding of DNA computations, and protein or peptide detection.
Researchers from the Tokyo University of Agriculture and Technology (TUAT) in Japan designed a beta-hairpin peptide (dubbed SV28) and formed a beta-barrel structure with several different sizes of nanopore ranging from 1.7 to 6.3 nm in diameter. The structure has three different regions of amino acid sequence: a beta-strand backbone containing 10 amino acids, a beta-turn with four amino acids, and two terminal structures. To verify the structure, the researchers performed molecular dynamics simulations, linear and exponential dependencies on the concentration, and applied voltage for the detection of double-stranded DNA.
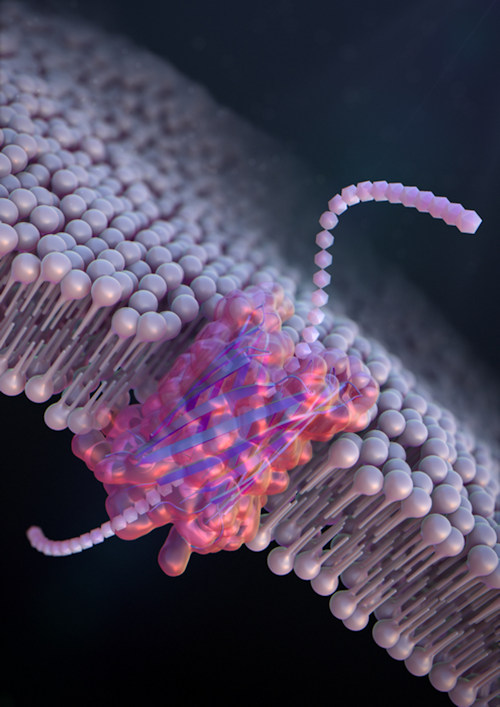
"Nanopore sensing is a powerful tool for label-free, single-molecule detection," said corresponding author Ryuji Kawano, PhD, a professor in the biotechnology and life science department at TUAT. "This is the first time that DNA and polypeptides were sensed using a de novo-designed nanopore."
The team also redesigned the beta-hairpin peptide by introducing a glycine kink that increases the curvature of the beta-barrel structure, resulting in stabilization of the overall conformation. The size of the SVG28 nanopore was estimated to be around 1.3 nm using molecular dynamics simulations. Detection of polypeptide chains was determined to be sensitive to both voltage and concentration, supporting the argument that the synthetic peptide could be used as a nanopore sensor.
"The folded structure of proteins is determined by their linear polypeptide sequence and gives rise to specific protein functionality," Kawano said, noting that all proteins have a unique structure and size. "The unique primary structure is the result of structural evolution such as the mutation and selection of amino acid residues over time. To reveal the relationship between this primary information and protein structure is one of the ultimate goals of science."
Next, Kawano's team plans to design various peptides and proteins to construct different types of nanopores to aid in peptide sequencing, operate as molecular robots, and more.
Do you have a unique perspective on your research related to synthetic biology? Contact the editor today to learn more.
Copyright © 2021 scienceboard.net