October 5, 2021 -- Scientists have developed a new visualization method to construct 4D models of X chromosome inactivation, lending insight into gene expression and opening new pathways to drug treatments for gene-based disorders and diseases. The findings are being published on October 8 in the Proceedings of the National Academy of Sciences of the United States of America.
X chromosome inactivation (XCI) is the process that turns off one X chromosome in female mammal embryos to ensure that females, like males, have only one functional copy of the X chromosome in each cell. XCI occurs when X inactive-specific transcript (Xist) RNA, a noncoding RNA on the X chromosome, spreads to coat the whole X chromosome and recruits various protein complexes to silence it.
A new model, 4DHiC, runs on supercomputers at the Los Alamos National Laboratory in New Mexico. It is named "4D" because it extracts 3D information from 2D XCI datasets and then renders the activity across time, which is considered the fourth dimension. The "HiC" part of the name refers to the Hi-C technique that enables large-scale sequencing of pairwise genomic contacts. The new approach incorporates existing 2D experimental Hi-C data from mice into its 3D model of the X chromosome.
"This is the first time we've been able to model all the RNA spreading around the chromosome and shutting it down," said first author Anna Lappala, PhD, a visiting scientist at Los Alamos and a polymer physicist at Massachusetts General Hospital and the Harvard Department of Molecular Biology, in a statement. "From experimental data alone, which is 2D and static, you don't have the resolution to see a whole chromosome at this level of detail."
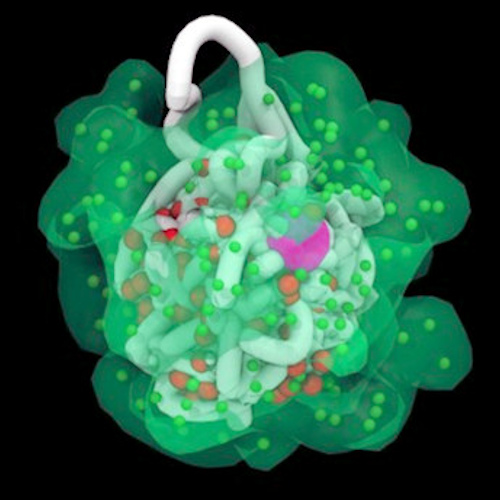
The visualization, which the authors posted on YouTube, shows Xist RNA particles, represented as small blue dots, swarming over the X chromosome, which appears as a tangled ball of spaghettilike strands that wiggle and bend as the X chromosome folds into a new shape, origami style. A large purple sphere at the center of the X chromosome represents the X inactivation center. The blue dots engulf and penetrate the depths of the chromosome, turning it off.
Answering questions about XCI
Previous work on XCI revealed specific structural units developing during the "folding" process of the X chromosome as it transitions from its active to inactive form. These structures include A/B compartments, which delineate active and inactive regions; S1/S2 compartments, which appear during the transition and have been assumed to be transitory; and a pair of megadomains, which are the structural end points of the folding process.
The 4DHiC model was able to reveal these structures in much more detail and clear up some remaining mysteries about them. For example, the model revealed how the A and B compartments merge to create S1 and S2 compartments, an intermediate and transitory state prior to the final superstructure of the inactivated X chromosome. Contrary to the prevailing belief that A/B organization is destroyed during XCI, the modeling showed that elements of these original structures persisted but remain hidden from view.
Like Google Earth for epigenetic processes
The model is a way for those not computationally savvy working in the field of epigenetics to jump into a chromosome and interact with it, according to co-author Dr. Jeannie Lee, PhD, professor of genetics at Harvard Medical School and vice chair of molecular biology at Massachusetts General Hospital.
She described their new modeling method as Google Earth for epigenetic processes, allowing anyone to "zoom into any location on an X chromosome, pick your favorite gene, see the other genes around it, and see how they interact," she explained.
Genetic and therapeutic applications
Knowing the structure of the X chromosome helps explain how genes are turned on and off. The new approach is expected to open new pathways to drug treatments for gene-based disorders and diseases.
"Our main goal was to see the chromosome change its shape and to see gene-expression levels over time," said Karissa Sanbonmatsu, PhD, a structural biologist at Los Alamos National Laboratory and corresponding author of the paper. "The hypothesis is that a compacted, tightly structured chromosome tends to turn off genes, but there are not a lot of smoking guns about this. By modeling 3D structures in motion, we can get closer to the relationship between structural compaction and turning off genes."
Variations in XCI itself can trigger a variety of developmental disorders. In addition, an inactive X can activate later in a process called age-related activation of inactive X. "It's associated with problems in blood cells in particular that are known to cause autoimmunity," Lee said.
Furthermore, the approach could lead to epigenetic treatments for mutations that cause disease. For example, certain neurological disorders in children can be treated by reactivating an inactive X chromosome to give the patient back something that's missing on their active X chromosome. "We think if we can reactivate the normal copy on the inactive X, then we would have an epigenetic treatment for that mutation," Lee said.
Do you have a unique perspective on your research related to epigenetics? Contact the editor today to learn more.
Copyright © 2021 scienceboard.net






