July 30, 2021 -- Advanced electron microscopy has enabled researchers to visualize how human viruses move in high resolution in a near-native environment. The visualization technique, published in Advanced Materials on July 24, could lead to an improved understanding of the molecular mechanisms of vaccines and therapeutics.
Cryogenic electron microscopy (cryo-EM) has become the gold standard for observing samples at or beyond atomic resolution. The technique requires flash freezing a sample and focusing a beam of electrons through it. The electrons interact with the sample's components, and cameras embedded in the instrument capture the deflected beams, resulting in images of individual molecules. Thousands of images can be processed to calculate what the item looks like in 3D. But to understand how the item functions in a natural setting, more images are needed.
"While cryo-EM can tell us a lot of information, it still produces a static image," said first author GM Jonaid, a doctoral student in the bioinformatics and genomics graduate program at the Huck Institutes of the Life Sciences, in a statement. "With improved chips and a powerful direct detector on the microscope, we can accumulate a lot of movie frames to view how the sample acts in real time. We can see things how they exist -- not just how we prepared them."
The research team, led by Deb Kelly, PhD, a professor of biomedical engineering at Penn State University, used liquid-phase electron microscopy (LP-EM) to study the 3D features of viruses in real-time. The team used high-frame-rate direct detectors, motion-correcting algorithms, and advanced computing procedures to generate 20-second videos of the viruses in motion.
Testing the power of LP-EM with adeno-associated viruses
Adeno-associated viruses (AAVs) are natural nanoparticles that can deliver vaccines or treatments directly to target cells. The virus's ability to enter human cells makes it an attractive system for delivering engineered payloads.
"AAV is a well-known gene therapy vehicle with current applications involved in drug delivery and vaccine development for COVID-19," said Kelly. "This model system is already well studied, so we can use it to validate our approach, with the goal of seeing biological entities in a liquid state as maintained in the human body."
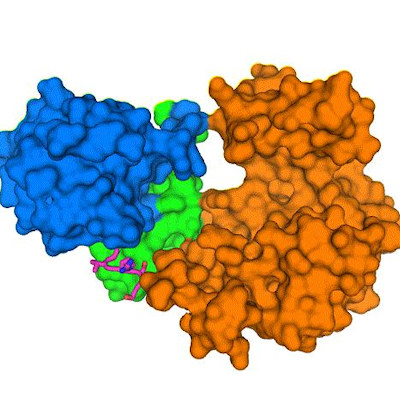
The team used silicon nitride microchips with integrated microwells to hold virus particles in liquid droplets for bright field transmission electron microscopy (TEM). The images showed the viruses moving slowly or diffusing in liquid. After processing the images, the researchers compared the AAV to a reference molecule and found that the test virus had subtle yet distinct structures, such as a lack of some N-terminal amino acids, which the authors suggested is needed to evade immune proteins. The researchers found that the AAV undergoes dynamic changes in a liquid environment, which are likely required to survive circulating in the blood and other tissues. The resolution of the images was approximately 3 to 4 angstroms (a single atom is measured as 1 angstrom).
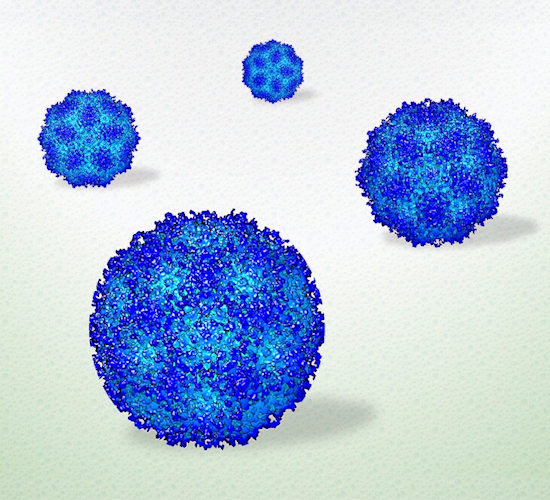
"The images are very comparable to cryo-EM data, but the preparation was less complex, less technically involved," said Jonaid. "Once we had the images, taken rapidly, like frames of a movie, we processed them just like we would any other high-resolution data."
The new method allowed the team to record real-time movies of viral assemblies for extended periods of time.
Testing relevant viruses: SARS-CoV-2
Once the researchers validated the methodology, they moved to a clinically relevant, albeit smaller target. They used the same imaging methods and computing algorithms to assess human immunoglobulin antibodies from the serum of COVID-19 patients.
"We saw how antibodies contained in the serum of COVID-19 patients interacted with the remaining SARS-CoV-2 particles," Kelly said, noting that the ability to observe such interactions would be especially useful when assessing the viability of vaccine candidates prior to clinical trials.
"Our results show new structures and active insights of human viruses contained in minute volumes of liquid -- the same size as respiratory droplets that spread SARS-CoV-2," Kelly continued.
The Penn State team plans to continue investigating the molecular underpinnings of SARS-CoV-2 and host receptor proteins using LP-EM as a complement to the information gathered from cryo-EM results.
"You really need data from both techniques to understand how viruses look and behave in the living body," Kelly said. "Visualizing the dynamic movement in solution complements high-resolution snapshots to reveal more complete information."
Do you have a unique perspective on your research related to bioimaging or virology? Contact the editor today to learn more.
Copyright © 2021 scienceboard.net