June 10, 2021 -- Scientists have untangled the function of proteins that block normal DNA repair processes and may permit unwanted cancer cell growth. New research published in Molecular Cell on June 7 details how the XRCC1 molecular scaffold protein works in tandem with the poly(ADP-ribose) polymerase (PARP) DNA break-detection protein to ensure that genetic errors are efficiently corrected.
DNA damage -- including nicks, scratches, and breaks -- is caused by normal cellular metabolic processes. The body has natural DNA repair processes that actively respond to fix DNA damage and keep cells functioning normally.
However, if DNA damage accumulates, errors in transcription can occur -- with fatal consequences. Many cancers have genetic mutations that cause DNA repair to be less effective than normal.
DNA base excision repair, a type of repair that causes only a single-strand break, is a highly conserved pathway in all organisms and is required for repair of DNA base damage. Beyond the core components of the system, mammalian cells employ additional proteins to accelerate the process, such as PARP1, PARP2, and XRCC1.
PARP proteins detect and are activated by DNA strand breaks, resulting in post-translational modifications that can help modify chromatin structure. PARP proteins can also recruit other proteins, such as XRCC1, to accelerate repair of DNA strand breaks. The role of XRCC1 during DNA base excision repair is not entirely clear because the type of repair it induces is only seen in a subset of events.
Now, a team of scientists at Tokyo Metropolitan University, the University of Sussex, and Kyoto University have revealed exactly how XRCC1 works with PARP proteins.
PARP1 activity resulting from elevated levels of unrepaired DNA breaks can result in cell death via nicotinamide adenine dinucleotide depletion, defective glycolysis, and necrosis/parthanatos (PARP-1 dependent cell death). In this way, excessive PARP1 blocks enzyme access and repair of single-strand break repairs. This discovery led to the development of anticancer drugs known as PARP inhibitors that work to disrupt the growth of certain types of tumors that depend on PARP proteins.
"PARP turns out to be something of a villain," explained Kouji Hirota, PhD, professor in the department of chemistry at Tokyo Metropolitan University. "The spots it marks become 'PARP traps,' which left unrepaired lead to dysfunction and cell death."
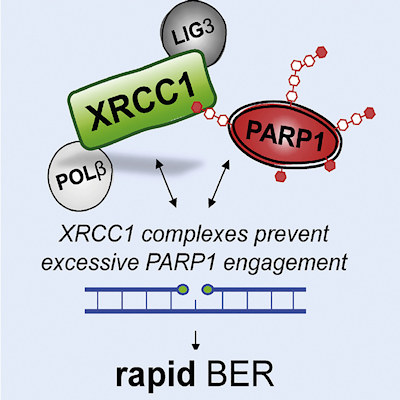
The scientists conducted in vitro cell culture experiments to compare cells lacking the XRCC1 gene with those lacking PARP or those lacking both proteins. The new work demonstrates that the essential role of XRCC1 during base excision repair is to prevent excessive PARP1 engagement and activity.
The team demonstrated that the accumulation of single-strand breaks and the cellular hypersensitivity to DNA base damage when XRCC1 is absent was eliminated with the depletion of PARP1. Without XRCC1 on patrol, PARP traps accumulate like landmines.
The authors proposed a model whereby PARP1 tends to associate excessively with repair intermediates, blocking their access and repair by other DNA base excision repair enzymes. The authors define this as a form of PARP1 trapping. When nicotinamide adenine dinucleotide becomes sufficiently depleted by excessive cycles of PARP1 association, the ability of PARP1 to dissociate from base excision repair intermediates is significantly reduced, leading to even tighter trapping and accumulation of PARP1 in chromatin.
Importantly, XRCC1 can assemble DNA polymerase β (POLβ) and DNA ligase III (LIG3) into protein complexes, which limits the number of opportunities for excessive PARP1 engagement and suppresses PARP1 trapping, ultimately promoting repair.
"PARP exerts toxic effects in the cell and XRCC1 suppresses this toxicity," Hirota elaborated.
In the future, the team seeks to delve even further into these processes, aiming to advance the development of future cancer treatments.
"These results indicate that XRCC1 is a critical factor in the resolution of PARP traps and may be a determinant of the therapeutic effect of PARP inhibitors used in the treatment of hereditary breast and ovarian cancer syndromes," said co-author Shunichi Takeda of Kyoto University.
Do you have a unique perspective on your research related to cancer research or molecular biology? Contact the editor today to learn more.
Copyright © 2021 scienceboard.net