May 27, 2021 -- Single-cell transcriptomics have been used to map which genes are turned on and off during the entire transmission cycle of the deadly human malaria parasite, Plasmodium falciparum. The results, published in Nature Communications on May 27, may lead to the identification of new gene targets for antimalarial treatments.
P. falciparum is a unicellular protozoan parasite and the deadliest cause of malaria in humans worldwide. The parasite is transmitted through the bite of a female Anopheles mosquito and causes falciparum malaria. Humans are the intermediate hosts of the parasite where asexual reproduction occurs, and the mosquitos are the hosts that support sexual reproduction.
P. falciparum develops in the midgut of mosquitoes before immature sporozoites travel to the salivary glands where they undergo further development and become infectious to humans. During the early phase, the parasite undergoes many stages of development.
Recently, the Wellcome Sanger Institute launched the Malaria Cell Atlas project as a data resource to provide scientists with a way to explore patterns of gene expression in individual parasites across multiple developmental stages and parasite species. However, for P. falciparum the atlas gave an incomplete picture of the parasite's entire developmental lifecycle.
A team from the lab of Jake Baum, PhD, at Imperial College London and the lab of Mara Lawniczak, PhD, from the Wellcome Sanger Institute have examined the development of P. falciparum in unprecedented detail. The team's work effectively adds a chapter to the Malaria Cell Atlas by completing the single-cell RNA sequencing (scRNA-seq) survey of the transmission stages from sexual forms transmitted to the Anopheles mosquito to the invasive sporozoite delivered to the human host.
The researchers isolated the different forms of the parasite and analyzed the activity of genes throughout the process using scRNA-seq. The final dataset included 1,467 single-cell transcriptomes spanning the broad developmental transitions from host to vector and vector to host transmission. These results were integrated with the existing 161 single-cell transcriptomes of mixed P. falciparum asexual stages, which enabled them to reconstitute the life cycle almost entirely.
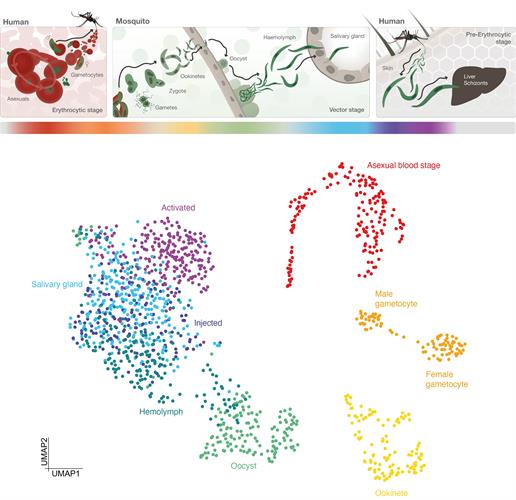
The results showed that cells from the same stage grouped together in 2D space and formed distinct cell clusters. The data also helped the team identify developmentally regulated transcripts that are expressed at highly variable levels across cells of the same stage. The authors suggested that this could theoretically account for differential phenotypes within a population and help predict if a particular parasite will be successful or not.
Understanding how each stage of development varies at the cellular level can help the scientists identify new targets that could be blocked to stop development and prevent transmission of the parasite.
"Being directly based on the human-infective parasite, our new data have clear implications for malaria control, which has an increasing focus on transmission blocking strategies both in terms of drugs that kill the parasite as it moves between stages and protective vaccines," said Eliana Real, PhD, postdoctoral scientist at the Imperial College London, in a statement. "Understanding how parasites behave transcriptionally within the mosquito vector provides a foundation from which new strategies will surely arise."
The scientists were interested in understanding how sporozoites infect humans and could provide a valuable target for antimalarial therapies and vaccine candidates. They were able to analyze transcriptional changes associated with development of the sporozoites over time to determine which genes are turned on during each discrete phase. This is the stage where the parasite is released into the human skin during a mosquito bite. The team was able to identify specific patterns of gene expression that define this critical stage of transmission.
"This fine granularity enables us to trace sporozoite developmental processes and to propose new mechanistic targets essential for each step and future vaccine targets for blocking malaria infection," said Virginia Howick, PhD, previously from the Wellcome Sanger Institute and now based at the University of Glasgow.
The team was also able to compare their dataset with a similar set from the related parasite Plasmodium berghei, a rodent malaria parasite that is often used as a model for studying malaria disease in the lab. This comparison showed which genes are conserved between species, and which are specific to the human version of the parasite.
"This level of gene surveillance at the individual parasite level throughout its life cycle will provide an invaluable resource for researchers to discover previously unexplored elements of Plasmodium cell biology, comparative Plasmodium species biology and the development of control methods that target particular pathways or lay the foundations for improving vaccines," commented Farah Dahalan, PhD, another postdoctoral scientist in the Baum Lab at Imperial.
Do you have a unique perspective on your research related to infectious diseases? Contact the editor today to learn more.
Copyright © 2021 scienceboard.net







