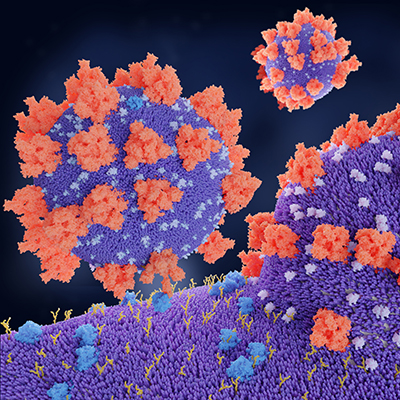
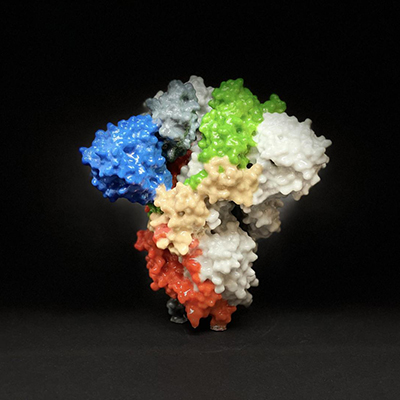
February 8, 2023 -- The infamous spike proteins on the surface of SARS-CoV-2 enable it to bind with and enter human cells. Because of their key role in spreading infection, these spike proteins are one of the main targets for COVID-19 vaccines and treatments.
Research from China, published February 8 in ACS Central Science, reports the discovery of small molecules that successfully target and bind with SARS-CoV-2 spike protein segments, locking them in a harmless closed configuration instead of the infectious open configuration.
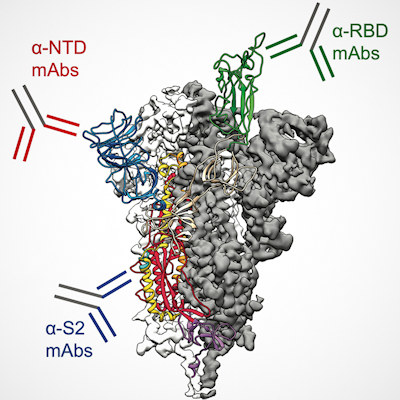
Spike proteins change shape when they attack a healthy cell. In their open structure, they expose a section known as the receptor-binding domain which attaches to the ACE2 protein on human cells. In the closed structure, this receptor-binding segment is tucked inside the spike protein and can't bind to human cells. Antibodies contained in some COVID-19 therapies or stimulated by vaccines or infection target the receptor-binding domain so it can no longer bind to ACE2.
However, because some emerging coronavirus variants contain mutations in the spike proteins' receptor-binding fragment, vaccines and antibody therapies designed to target that fragment may become less effective as the virus mutates.
Targeting other less mutation-prone parts of the spike protein is one alternative. One pocket in the spike protein has been dubbed the Achilles' heel of the virus. When this pocket is occupied by free fatty acids or other compounds, the protein remains locked in its closed, harmless configuration. However, because those compounds are unstable or bind weakly, they aren't considered suitable treatments.
The researchers sought other potential treatments that lacked these flaws. Using computer modeling, the team screened a library of small molecules, seeking those that could slip into this pocket and bind firmly to the spike protein, keeping it in the closed position. The researchers then used various techniques, including surface plasmon resonance, a reflectivity measurement used to detect molecular adsorption, to evaluate analogs of these molecules for improved binding and solubility.
The researchers contend that the resulting compounds, which can bind to spike proteins from the original coronavirus as well as the omicron BA.4 variant, could serve as a starting point for the development of broad-spectrum COVID-19 treatments.
Copyright © 2023 scienceboard.net