October 13, 2020 -- In order to determine optimal cell growth conditions for the development of cellular therapies, researchers have devised an algorithm called EpiMorgify that can predict what factors are needed to produce high-quality cultures. The study was published on October 9 in Cell Systems.
Typically, cell cultures use an undefined culture medium containing many different components or a standard medium to which is added certain defined sets of cell-type-specific factors such as growth factors or extracellular matrix components. These factors are often determined by domain knowledge and/or trial and error.
This methodology does not allow for chemically defined media (growth media for which all chemical components are known) and directed differentiation (differentiation in vitro toward a specific cell or tissue type), which are often required for predictable manufacturing of cell therapies.
The researchers at Duke-NUS Medical School, Singapore, and Monash University in Australia have developed a computational tool that can systematically identify chemically defined maintenance and conversion cell culture conditions.
While every cell type has the same genome, the specialized nature of an individual cell type is derived from differences in its epigenetic, transcriptional, and protein landscape. For example, the H3K4me3 histone modification is known to be associated with active genes, and as such is a key epigenetic regulator.
Conversely, H3K27me3 histone modifications are known to silence nonlineage-specific genes but are also important developmental regulators. Using this knowledge, the researchers modeled a cell's epigenetic state to identify key drivers of cell identity.
The approach, called EpiMorgify, uses available epigenetic datasets and models a cell's epigenetic state using H3K4me3 and H327me3 as filters. Using thresholds and incorporating protein-protein interaction network information to prioritize genes that exert large regulatory effects over cell identity, the model can predict components for maintenance and conversion conditions in many cell types.
In the new study, the model was used to rank receptor-ligand pairs and add those that appear to have a strong regulatory influence to the culture media.
"This approach facilitates the identification of the optimum cell culture conditions for converting cells and also for growing the high-quality cells required for cell therapy applications," said senior author Jose Polo, PhD, from Monash University's Biomedicine Discovery Institute and the Australian Research Medicine Institute, in a statement.
To predict factors that facilitate cellular conversion (differentiation), the researchers used EpiMorgify to compute the change per gene between source cells and target cell types, which was associated with regulatory network information. The model was then used to identify receptor-ligand pairs that are required for conversion.
"This study leverages our expertise in computational and systems biology to facilitate the good manufacturing practice (GMP) production of high-quality cells for these much-needed therapeutic applications," said senior author Enrico Petretto, PhD, associate professor at Duke-NUS.
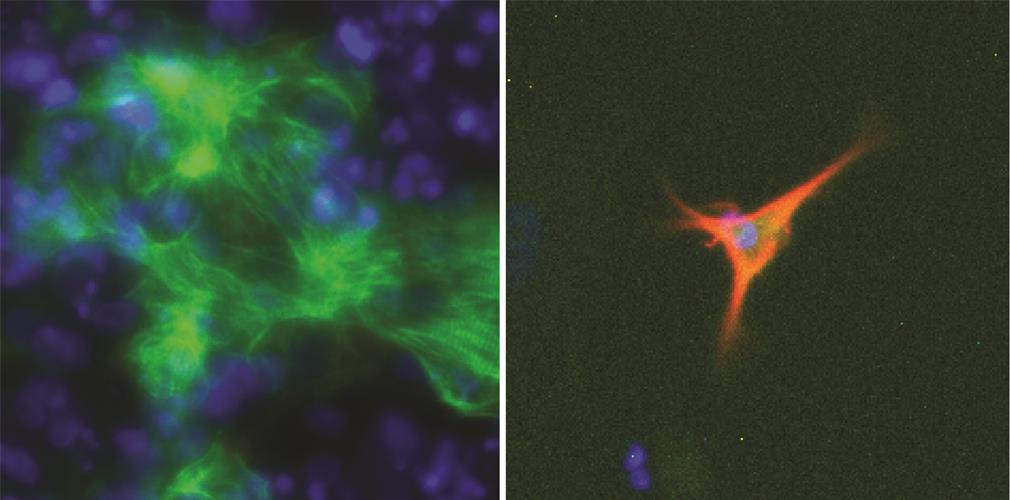
The researchers validated EpiMogrify by predicting which factors are required for growth of astrocytes and cardiomyocytes and maintaining the cells in vitro. The model was also able to predict factors that promote the generation of astrocytes and cardiomyocytes from neural progenitors and embryonic stem cells. EpiMorgify accurately predicted fibronectin at first rank ligand for astrocytes and cardiomyocytes, which is commonly added to undefined matrix during culture.
The predictions from the models were compared to existing undefined conditions. In both cases, the predictions performed as well or better than the undefined conditions, displaying increased cell growth and survival and higher differentiation efficiency.
"The developed technology can identify cell culture conditions required to manipulate cell fate and this facilitates growing important cells in chemically-defined cultures for cell therapy applications," added lead author Uma S. Kamaraj, PhD, graduate of Duke-NUS Medical School.
The researchers have filed a patent for their approach and the cell culture factors that it predicted are tied to cell fate. EpiMogrify's predicted molecules are available for other researchers to explore on a public database: http://epimogrify.ddnetbio.com.
Do you have a unique perspective on your research related to cell culture or cell biology? Contact the editor today to learn more.
Copyright © 2020 scienceboard.net