July 31, 2020 -- The crystal structure of nonstructural protein 16 (nsp16) of SARS-CoV-2, which plays a role in viral RNA capping to mimic host messenger RNA (mRNA), reveals specific rational design targets that could be used to develop effective therapies against SARS-CoV-2 and other coronaviruses. The research was published in Nature Communications on July 24.
The SARS-CoV-2 genome encodes four structural and 16 nonstructural proteins that are essential for the lifecycle of the virus. It exploits the host cell environment for its own replication, but to achieve this, viral RNA must be protected from the host innate immune system. Furthermore, RNA located in the cytoplasm, like viral RNA, is prone to degradation.
Viral RNA capping to mimic host mRNA
As recently shown, RNA capping that resembles host mRNA protects the integrity of the viral RNA and ensures translation of viral proteins. The virus possesses its own cap-synthesizing enzymes including nonstructural proteins nsp10, 13, 14, and 16. Nsp13 primarily functions to unwind viral RNA during replication and can therefore be considered a helicase.
Nsp14 and nsp16 are responsible for the methylation of the cap and are both S-adenosylmethionine (SAM)-dependent methyltransfereases (Mtase). The enzymatic activity of Mtase is enhanced by nsp10, which acts as a cofactor.
A type 1 RNA cap follows the following process: a phosphate is removed by 5'-RNA triphosphatase, a guanosine monophosphate (GMP) is added to the end of RNA by guanylyltransferase (GTP), followed by two methylations (N-7 methylation of the first nucleobase by nsp14 and C2'-oxygen methylation of the next nucleotide by nsp16).
Informing structure-based drug design targeting nsp16
Nsp16, a 2'-oxygen Mtase, is a promising molecular drug target that is indispensable for replication of coronaviruses. Researchers from the Institute of Organic Chemistry and Biochemistry (IOCB) of the Czech Academy of Sciences determined the x-ray crystal structure of SARS-CoV-2 nsp10-nsp16 in complex with sinefungin, a pan-Mtase inhibitor.
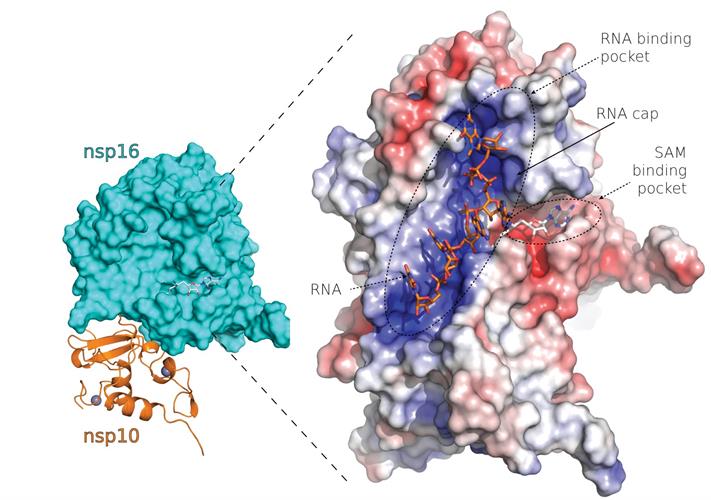
The researchers confirmed an overall fold similarity between SARS-CoV and SARS-CoV-2 nsp10-nsp16 complexes and a high conservation of the SAM-binding site among coronaviruses. There are six residues localized in a canyon within the nsp16-binding site that are absolutely conserved among SARS-CoV-2, SARS-CoV, and MERS-CoV, highlighting the importance of RNA methylation for coronaviruses.
The structure revealed a mixed alpha-beta fold with the Mtase inhibitor bound in a central canyon. Nsp10 and nsp16 interact through a large network of hydrogen bonds mediated by water molecules or hydrophobic interactions.
There is currently no unliganded coronavirus nsp16 protein structure available, so the mechanism of activation is not confirmed. However, nsp10 showed no conformation change upon binding to nsp16, so it is expected that nsp10 induces a conformational change in nsp16 Mtase that switches nsp16 into an active enzyme.
The team found several factors that could be used in therapeutic design. First, the nsp10-nsp16 complex must be able to bind methylated GTP and substrate for the methylation reaction (SAM) within a canyon of nsp16. Second, rational structure design must include an inhibitor that comes into direct contact with the 2'-hydroxyl group of the first nucleotide.
Do you have a unique perspective on your research related to drug design or virology? Contact the editor today to learn more.
Copyright © 2020 scienceboard.net


