December 16, 2019 -- Researchers from the University of California San Diego have developed a brand new CRISPR-based gene-drive system that dramatically increases the efficiency of inactivation of genes responsible for antibiotic resistance. The new system is detailed on December 16 in Nature Communications.
Genes conferring antibiotic resistance are often found on plasmids, circular forms of DNA that can replicate independently of the bacterial genome. Amplification effects of these plasmids can lead to the transfer of antibiotic resistance among bacteria. This poses a significant challenge to disrupt this function. Researchers have developed several cut-and-destroy methods but have had only moderate success with them.
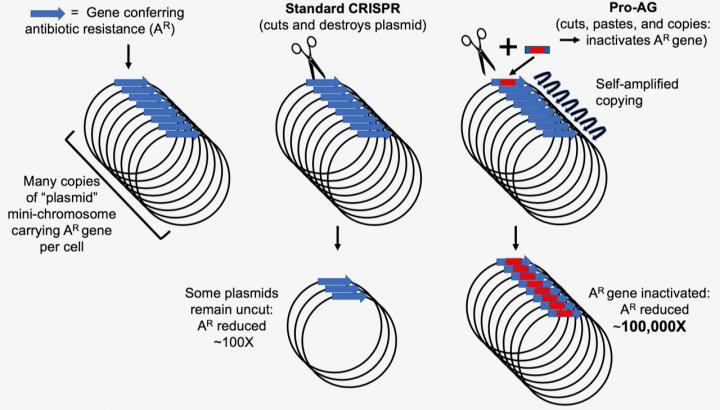
Plasmid-encoded antibiotic resistance determinants can be targeted by CRISPR gRNA/Cas9 systems in bacteria. A new CRISPR tool could be instrumental in preventing pathogenesis of human bacterial infections and plasmid-borne antibiotic resistance genes and virulence determinants. To achieve this system, researchers developed a proactive genetic system, or ProAG, that uses self-copying mechanisms to scrub antibiotic resistance.
The new CRISPR-based gene-drive system uses guide RNA (gRNA) to insert linked cargo sequences to the Cas9-mediated cleavage site. The functional gRNA cassette containing the Pro-AG sequences inactivates an antibiotic resistance marker (bla) on a high copy number plasmid. The system relies on a self-amplifying "editing" mechanism that increases its efficiency through a positive feedback loop. The success of this system is attributed to the precise insertion of the gRNA into the targeted cleavage site with appropriate cargo sequences, which reduced antibiotic-resistant colonies.
While Pro-AG is not yet ready for treating patients, "a human delivery system carrying Pro-AG could be deployed to address conditions such as cystic fibrosis, chronic urinary infections, tuberculosis and infections associated with resistant biofilms that pose difficult challenges in hospital settings," said Victor Nizet, distinguished professor of Pediatrics and Pharmacy and the faculty lead of the UC San Diego Collaborative to Halt Antibiotic-Resistant Microbes (CHARM).
Moreover, the system can edit large plasmids or single-copy genomic targets to introduce functional genes, thereby expanding the available toolkit for biotechnology and genetic engineering. Because Pro-AG "edits" its targets rather than destroys them, this system also enables engineering or manipulating bacteria, rendering them harmless or even recruiting them to perform beneficial functions.
"The highly efficient and precise nature of Pro-AG should permit a variety of practical applications, including dissemination of this system throughout populations of bacteria using one of several existing delivery systems to greatly reduce the prevalence of antibiotic resistance in the environment," said Ethan Bier, a distinguished professor in the Section of Cell and Developmental Biology and science director of the UC San Diego unit of the Tata Institute for Genetics and Society (TIGS).
Do you have a unique perspective on your research related to genetic engineering? Contact the editor today to learn more.
Copyright © 2019 scienceboard.net


