February 5, 2020 -- A novel imaging technique improves the processing time of hyperspectral fluorescence imaging (fHSI) and allows scientists to peer more deeply and clearly into living organisms. Researchers from the University of Southern California (USC) Michelson Center for Convergent Bioscience have been developing the technology for years and published their findings in Nature Communications on February 5.
fHSI is a common technique for imaging multiple fluorescently tagged components in biological samples that allows scientists to follow cells of tissues in both space- and time-wise manners. It can differentiate colors across a spectrum to produce spectacularly vivid color images of cells and tissues.
Traditional imaging techniques differentiate fluorescent protein signals based on wavelength, but overlapping signals limit the number of fluorescent tags that can be used. fHSI overcomes this challenge allowing for an expanded fluorescent palette of different labels. However, with these colorful images, the complexity of datasets is increased, and computational processing is very time-consuming.
"There is a gap between acquisition and analysis of the hyperspectral data, where scientists and doctors are unaware of the information contained in the experiment. SEER is designed to fill this gap," said lead study author Francesco Cutrale, PhD, a research assistant professor of biomedical engineering at the USC Viterbi School of Engineering.
Preprocessing visualization can be beneficial and allows scientists to evaluate image collection parameters and choose the most appropriate processing method. The researchers developed an approach they called Spectrally Encoded Enhanced Representations (SEER), which was developed to provide improved and computationally efficient simultaneous color visualization of multiple spectral components of hyperspectral fluorescence images.
How it works
SEER relies on mathematical computations to parse fluorescence data faster. In fact, SEER can process a 3.7 GB dataset with 1.26 x 108 spectra in 6.6 sec, and a 43.88 GB dataset with 1.47 x 109 spectra in 87.3 sec. This is 67 times faster and at 2.7 times greater definition than present techniques.
The execution of SEER has a simple foundation. Spectrally representative color maps are generated based on phasors (phase vectors) to enhance the visualization of multi- and hyperspectral imaging. SEER transforms the spectrum at each pixel into its Fourier components, by means of a reference color map. Each point of the phasor plot can then be easily remapped to the original fluorescent image.
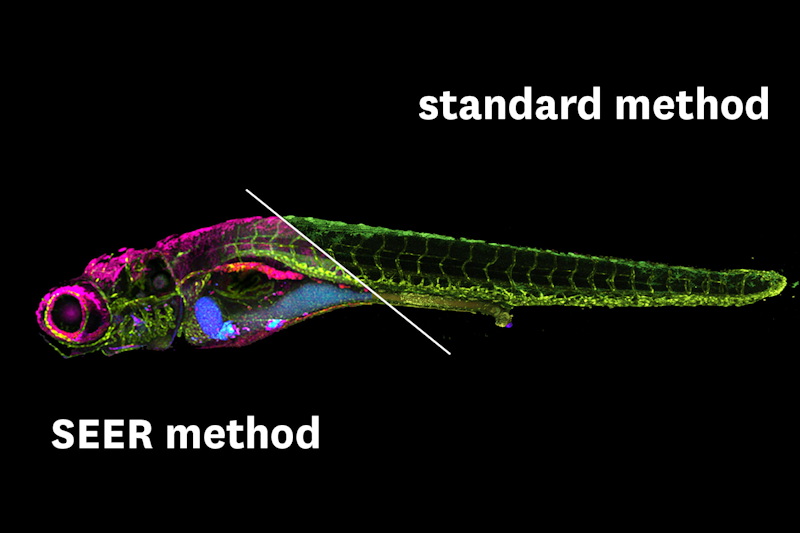
The results of SEER show an enhanced visualization of spectral properties, representing distinct fluorophores with distinguishable pseudo-colors and mathematically highlighted differences between intrinsic signals during live imaging. SEER can be applied quickly and efficiently even with large data sizes, circumventing the typical prolonged and expensive hyperspectral processing. The system is also flexible in nature. The users can apply several different standard reference maps to determine which is more appropriate for their data and enhance the most important image features.
SEER can be applied to fluorescence or to standard reflectance hyper- and multispectral imaging. Live-imaging visualization is another potential application for SEER, as it minimizes photobleaching and phototoxicity.
One limitation of SEER is that it is currently intended as a preprocessing visualization tool and, currently, is not utilized for quantitative analysis.
SEER's first application will be in the medical and research field. The algorithm, first authored by researchers at the Translational Imaging Center of USC, will be used for detecting early stages of lung disease and potential damage from pollutants in patients in a collaboration with doctors at Children's Hospital Los Angeles. Also, scientists in the life sciences field have started adopting SEER in their experimental pipelines in an effort to further improve the efficiency of current imaging techniques.
Do you have a unique perspective on your research related to microscopy or disease research? Contact the editor today to learn more.
Copyright © 2020 scienceboard.net






