December 18, 2019 -- Researchers develop an algorithm that locates specific topologically associating domains (TADs) which are implicated in disease development, including cancer. The technology, called OnTAD, can examine internal architectures of TADs, which are important in elucidating their biological functions. The work was published in Genome Biology on December 18.
Within the genome there are local regions where interactions are significantly higher than adjacent regions, called topologically associating domains (TADs). These regions form in conserved locations between species, indicating that they are basic architectural units that play a role in regulatory interactions. Recent studies show that TADs contain internal substructures, but scientists still lack a comprehensive understanding of the functions of hierarchical structures within TADs.
Researchers from Pennsylvania State University discovered that disruption of TADs can expose genes to improper regulatory elements and can lead to aberrant gene expression, causing initiation of cancer, for example. Therefore, the team developed OnTAD, an optimized nested TAD caller that uncovers hierarchical TAD structures from high throughput sequencing-based method, Hi-C, data.
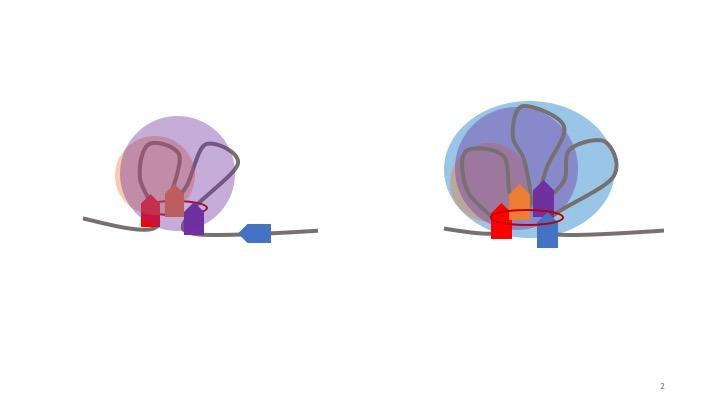
By studying the genomic and epigenetic profiles of specific TADs using OnTAD, researchers were able to make several important conclusions about the biological function of TADs. These more densely packed coils of DNA inside TADs have greater gene expression due to close proximity to their regulatory elements.
"Think about New York City, with its boroughs, neighborhoods within boroughs, and street locations within neighborhoods. Each level of organization is nested within a higher level," explained Ross Hardison, T. Ming Chu Professor of Biochemistry and Molecular Biology, Penn State. "Just like you are more likely to interact with someone on the same street rather than someone in another borough, DNA interactions are more frequent within the inner-most nested TADs. This is important because interactions among DNA segments -- such as genes and enhancers -- are needed for proper gene regulation. The OnTAD algorithm rapidly and efficiently reveals these levels of organization in DNA interactions."
They found that inner TAD hierarchies are more epigenetically active than outer TADs. They also found that TADs with hierarchical structures had more active epigenetic states (due to higher cohesion protein enrichment) than TADs without hierarchical structures. Epigenetic changes control important cell processes such as cell memory and identity.
"As we better understand how DNA interactions function in normal gene regulation, we can be better prepared to uncover how mutations in DNA can alter those interactions that can lead to incorrect gene expression and influence the development of cancers and other diseases," said Hardison.
"These results demonstrate that OnTAD is a powerful tool for revealing different levels of DNA organization across a genome," said Qunhua Li, associate professor of statistics, Penn State. "It should facilitate improved investigations into the roles of this organization in gene regulation."
Do you have a unique perspective on your research related to molecular biology? Contact the editor today to learn more.
Copyright © 2019 scienceboard.net






