November 10, 2022 -- Researchers have combined whole-genome sequencing with other techniques to generate detailed quantitative maps of genetic subclones in tumors, which they contend provide a complete view of cancer that has not been previously possible.
Genome sequencing has previously revealed mosaics of different subclones present in the same tumor without uncovering the exact spatial growth patterns and underlying mechanisms. However, in a paper published November 9 in the journal Nature, a team of researchers from multiple European research centers describe a push to address the need for ways to elucidate the growth patterns and mechanisms.
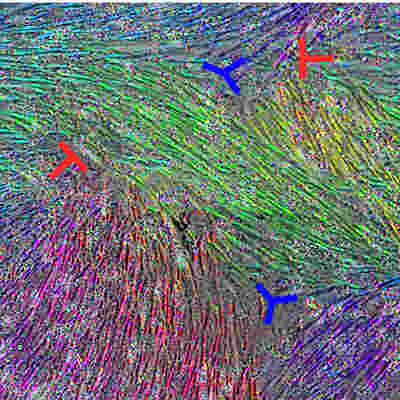
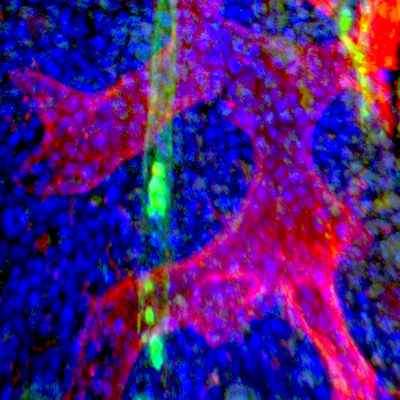
The new technology they developed taps hundreds of thousands of tiny fluorescent molecular probes to interrogate cellular DNA and RNA and scan large pieces of tissue using fluorescence microscopy. Leveraging these probes to target dozens of mutations in a set of cancer cell clones, the innovative technique allowed researchers to accurately reconstruct the spread of the clones.
Artem Lomakin, first author from EMBL's European Bioinformatics Institute and the German Cancer Research Center, described the system and its significance in a statement to publicize the paper.
"We have created a system that combines computational and experimental techniques that allows us to map evolutionary cancer lineages in their natural habitat of human tissue. While it has been previously possible to trace the lineage of cancer tumor cells in an experimental setup, this is the first time that multiple lineages were traced in human tissues, giving a complete overview of breast cancer development in the body. Insights generated by our system were impossible to get before, especially at this scale," Lomakin said.
Lomakin and collaborators from Wellcome Sanger Institute, the Science for Life Laboratory in Sweden, and his own employers used "whole-genome sequencing, followed by highly multiplexed base-specific in situ sequencing, single-cell resolved transcriptomics, and dedicated algorithms to link these layers" to address the challenge.
The resulting system generates maps that provide the basis for studying clonal growth patterns and the histological characteristics, microanatomy, and microenvironmental composition of each clone. Applied to tissue sections from primary breast cancers, the workflow revealed "intricate subclonal growth patterns."
Across multiple stages of breast cancer development there were specific patterns of clone growth. The behavior of the clones differed depending on their starting location in the breast, suggesting that factors other than genetics -- the current focus of cancer drug development and treatment -- influence how tumors grow.
"An important insight from our research is that it may not be the genetic changes alone that are the reason that the cancer cells survive and spread; it could also be where they are. This adds an additional layer of complexity as well as new potential ways to target the disease," said Mats Nilsson, co-senior author from the Science for Life Laboratory at Stockholm University.
In theory, the research suggests therapies could limit the growth of cancer influencing the environment around the tumor. The new workflow could also show how treatments affect both the cancer and its interaction with the immune system.
Copyright © 2022 scienceboard.net