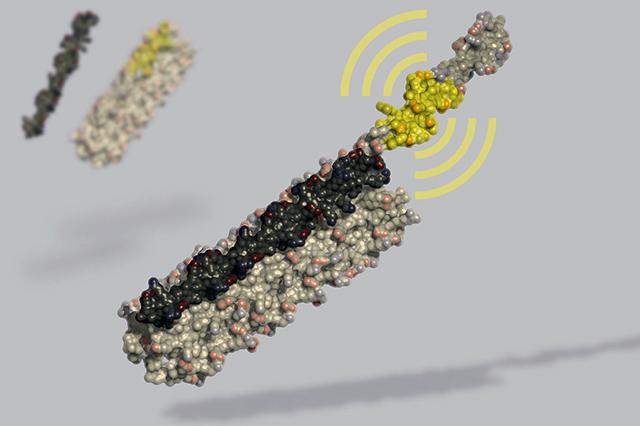
July 29, 2019 -- A new de novo designed protein – degronLOCKR, is the first completely artificial protein switch that operates within cells. LOCKR (short for Latching, Orthogonal Cage/Key pRotein) design and initial practical applications were published in Nature on July 24, 2019. Research was conducted by research teams at UW Medicine Institute for Protein Design and the University of California San Francisco.
This new protein acts as a sort of robot, performing certain tasks (modifying gene expression, redirecting cellular traffic, degrading other proteins, etc…) under specific conditions (biological circuits act as sensors to detect changes in internal and external cellular environments). This is the first biotechnology tools designed and built entirely by humans.
The research team used computational protein design to create self-assembling proteins that present bioactive peptides only upon addition of specific molecular "keys." LOCKR is designed as a five-helix ‘cage’ with a single interface that can interact either intramolecularly with a terminal ‘latch’ helix or intermolecularly with a peptide ‘key’. To test the design, researchers installed LOCKR in yeast and made it degrade a specific cellular protein at a time designated by the researchers.
"LOCKR opens a whole new realm of possibility for programming cells," said Ng, a graduate student at UCSF. "We are now limited more by our imagination and creativity rather than the proteins that nature has evolved."
Deep Learning and Proteins
Last year, Google debuted an algorithm called Alphafold that uses a two-step process to predict protein structure. Scientists are working towards using AI to predict protein arrangement and ultimately dictate protein function. Previously, scientists have only been able to determine protein structure by laboratory techniques such X-ray crystallography. Another scientist, Mohammed AlQuraishi, a biologist at Harvard Medical School in Boston, Massachusetts, has developed a new algorithm that predicts proteins in a single step and within milliseconds, utilizing a neural network based on the design of the human brain. He published his work in Cell Systems and hopes that other will build upon the foundation that he has set. These algorithms have competed in CASP13, the 13th Critical Assessment of Structure Prediction, a biennial competition aimed at predicting the 3D structure of proteins. Eventually, the computational predictions that arise will be powerful and accurate enough to be used in drug design and to better understand disease.
Designer proteins
Traditional methods for determining protein structure, like x-ray crystallography, are laborious and scientists cannot predict the shape from the sequence alone. Therefore, scientists have started designing artificial proteins with various functions. De novo protein engineering requires weeks of computational prediction time and months of iteration to determine viable candidates. Why? It’s all about the folding. The multiple forces holding proteins together restrict its shape and require precise angles and bonds to be present to accommodate the forces.
One lab from the University of Washington uses a molecular modelling and search tools called Rosetta, which can calculate the energy of a folded protein and search for the lowest energy sequence for a given structure, or the lowest energy structure for a given sequence. Rosetta was a contender in CASP13.
Once protein structure can be predicted, then researchers, can be manipulated to create new structures and shapes. This is where the LOCKR protein comes back into the picture. Complications occur when candidate proteins are put to the test when they must perform a specific function. Complications can include: misaligned structures, structures that do not bind tightly enough, or the placement of side chains is not in the correct position. This is why designing a de novo protein with functionality is truly groundbreaking.
What do you think will be the next breakthrough in protein design?
Do you have a unique prospective on your research related to protein design. The Science Advisory Board wants to highlight your research. Contact the editor today to learn more.
Copyright © 2019 scienceboard.net