May 4, 2021 -- NanoString Technologies' GeoMX digital spatial profiler (DSP) technology has been used to develop a foundational dataset to better understand the biological effect of SARS-CoV-2 infection in human lungs and support the identification of new therapeutic interventions and prevention strategies.
The research characterized autopsy lung tissue samples from patients who had been infected by SARS-CoV-2. It was published in Nature and conducted by a network of scientists from the Broad Institute and Beth Israel Deaconess Medical Center, Massachusetts General Hospital (MGH), the Ragon Institute of MGH, Massachusetts Institute of Technology, Harvard, Brigham and Women's Hospital, and Columbia University Irving Medical Center, in addition to scientists at NanoString.
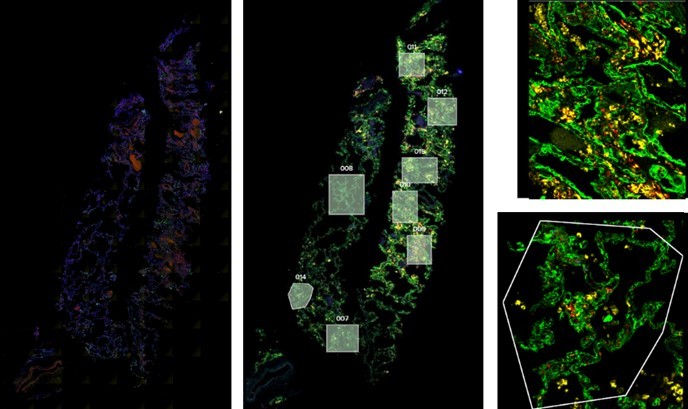
GeoMx DSP enables researchers to rapidly and quantitatively characterize tissue morphology with a high-throughput, high-plex RNA and protein profiling system.
In conjunction with single-cell RNA sequencing data, researchers utilized the cancer transcriptome atlas (CTA) and the whole transcriptome atlas (WTA), with additional custom content specific to SARS-CoV-2 genes, to produce and spatially map a comprehensive cell-type atlas. These data will help pinpoint cellular processes, expression pathways, and immune cell profiles to offer insights into systemic COVID-19 disease progression.
Copyright © 2021 scienceboard.net


