March 5, 2020 -- Researchers from the U.S. Department of Energy's Oak Ridge National Laboratory have used the Summit supercomputer to identify 77 small-molecule drug compounds that potentially target the SARS-CoV-2 virus, which is responsible for the novel coronavirus disease (COVID-19) outbreak.
The screen was conducted on over 8,000 compounds that are likely to bind to the spike protein of coronaviruses. The researchers ranked the results and published their findings on ChemRxiv, a preprint server for chemistry.
First, they designed computational models based on genetic sequences published in the literature. Next, they programmed Summit to run a chemical simulations code to perform molecular dynamics simulations to analyze the movement of atoms and particles in the spike protein.
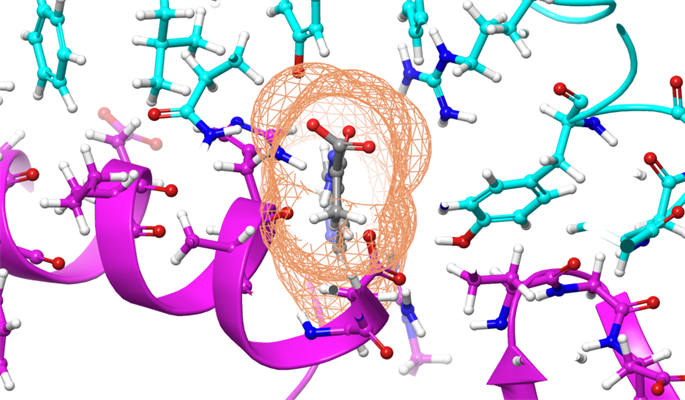
Summit generated a list of 77 small-molecule compounds, including medications and natural compounds, that may warrant further experimental testing. As updated models of the SARS-CoV-2 virus are published, the team plans to run the computational study again with new versions of the spike protein. The researchers emphasize that experimental studies are necessary to verify the efficacy of the identified chemicals against SARS-CoV-2.
"Summit was needed to rapidly get the simulation results we needed. It took us a day or two, whereas it would have taken months on a normal computer," said Jeremy C. Smith, PhD, governor's chair at the University of Tennessee (UT) and director of the UT/Oak Ridge National Laboratory Center for Molecular Biophysics. "Our results don't mean that we have found a cure or treatment for the Wuhan coronavirus. We are very hopeful, though, that our computational findings will both inform future studies and provide a framework that experimentalists will use to further investigate these compounds. Only then will we know whether any of them exhibit the characteristics needed to mitigate this virus."
Copyright © 2020 scienceboard.net


