January 24, 2020 -- Leveraging the power of new imaging techniques, scientists are using unique fluorescent tags in combination with superresolution microscopy to gain novel insights into a receptor that plays a key role in controlling blood sugar levels. These insights will open the door to improved treatment options for type 2 diabetes.
Glucagon-like peptide-1 receptor (GLP1R) is a class B G protein-coupled receptor (GPCR) involved in metabolism, specifically hormone regulation. The endogenous ligand, glucagon-like peptide-1 (GLP-1) is released in the gut in response to food intake, ultimately causing a cascade effect for the amplification of insulin release.
Current methods of visualizing GLP1R have a number of limitations, one of which is incompatibility with superresolution imaging techniques. In a new study, an international research team led by scientists from the University of Birmingham and the Max Planck Institute for Medical Research presented a new methodology to examine the receptors. The team generated a specific probe for endogenous GLP1R detection using LUXendins, far-red fluorescent antagonists. The technique was published on January 24 in Nature Communications.
The researchers used several techniques, including synthesis of marker compounds, immunostaining, superresolution microscopy, and in vivo examination of mice to locate GLP1R within cells and monitor its response to signal molecules. The results of the study provide an updated view of GLP1R expression patterns in pancreatic islets, brain, and β-like cells. The team was also able to use the technique to demonstrate that endogenous GLP1Rs form nanodomains at the membrane and to reveal receptor subpopulations with distinct diffusion modes in their nonstimulated state.
In the study, LUXendin645 allowed for multiplexing with primary monoclonal antibodies or SNAP self-labeling suicide enzymes. Further, LUXendin645 labeling was widespread in the islet and well colocalized with insulin immunostaining. The researchers verified the specificity of LUXendin645 by using GLP1R knock-out mice which showed intact response to glucose but absence of exendin-stimulated insulin secretion.
Superresolution imaging with LUXendin651 revealed higher-order GLP1R organization. They found that in beta cells GLP1R was not randomly arranged but rather tended to organize into nanodomains with neighbors.
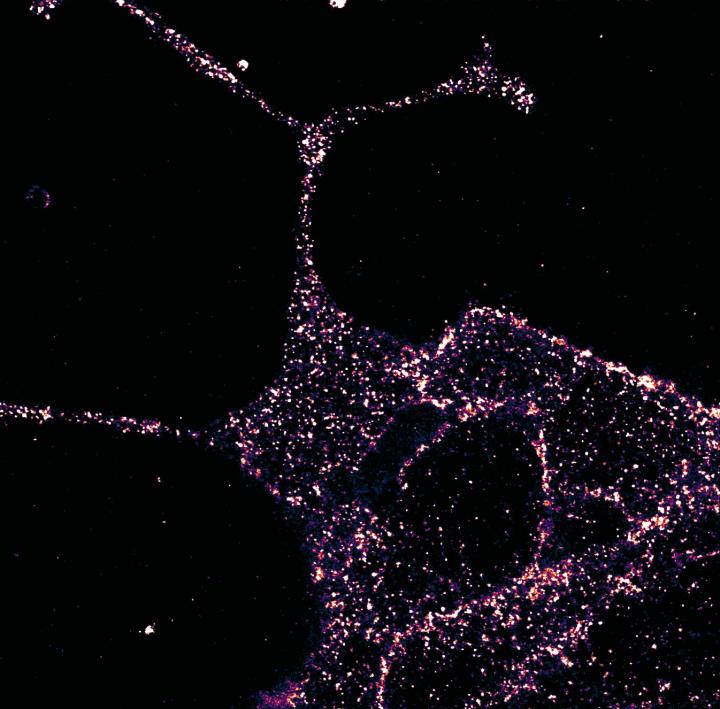
The researchers explored if LUXendins could be used to track single GLP1Rs in live cells with single-molecule microscopy (total internal reflection fluorescence microscopy). Using this technique with LUXendin651, they showed that GLP1R diffuses the membrane in a nonstimulated state. LUXendin555 allowed for in vivo labeling of islets via two-photon imaging.
"Our experiments, made possible by combining expertise in chemistry and cell biology, will improve our understanding of GLP1R in the pancreas and the brain," said Johannes Broichhagen, PhD, departmental group leader of the Max Planck Institute for Medical Research. "Our new tools have been used in stem cells and in the living animal to visualize this important receptor, and we provide the first superresolution characterization of a class B GPCR. Importantly, our results suggest a degree of complexity not readily appreciated with previous approaches."
Do you have a unique perspective on your research related to bioimaging or disease research? Contact the editor today to learn more.
Copyright © 2020 scienceboard.net


